Bioinformatic Analysis of Circadian Expression of Oncogenes and Tumor Suppressor Genes
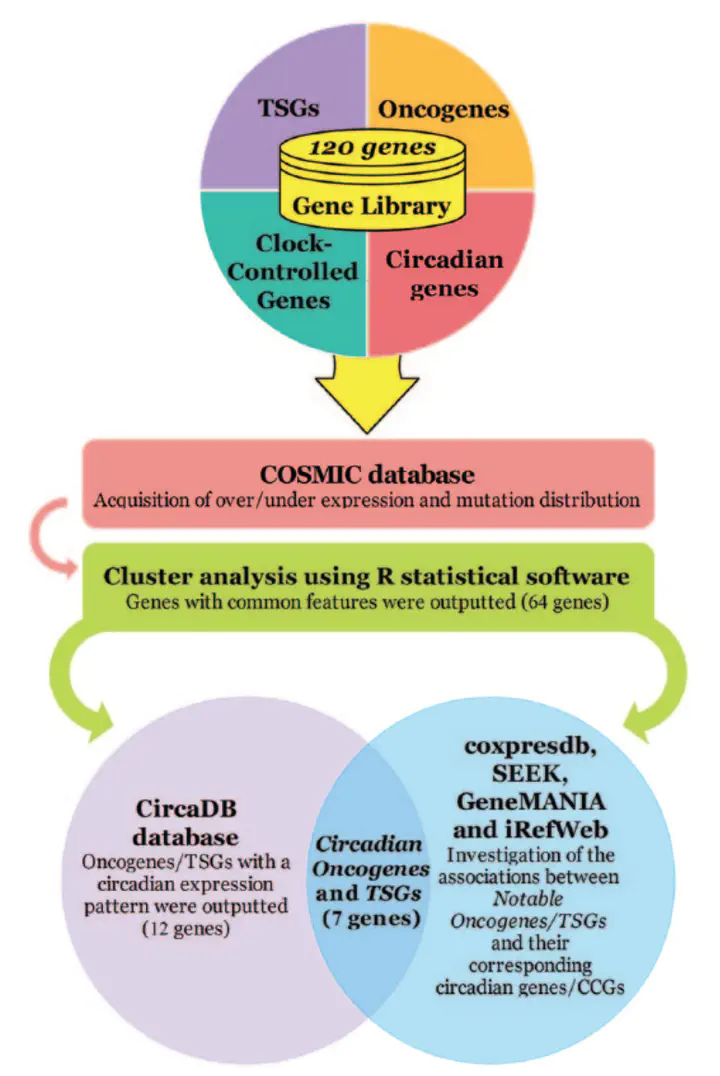 Image credit: Adrian Salavaty
Image credit: Adrian SalavatyAbstract
Methods: A bioinformatic analysis was conducted on a gene library comprising 120 genes to investigate the circadian expression of human oncogenes and TSGs. To achieve this purpose, the genotranscriptomic data were retrieved from COSMIC and analyzed by R statistical software. Furthermore, the acquired data were analyzed at the transcriptomic and proteomic levels using several publicly available databases. Also, the significance of all analyses was confirmed statistically.
Results: Altogether, our results indicated that 7 human oncogenes/TSGs may be expressed and function in a circadian manner. These oncogenes/TSGs showed a circadian expression pattern at CircaDB database and associated with at least one of the circadian genes/CCGs based on both genotranscriptomic and correlation analyses.
Conclusions: Although 4 of 7 finally outputted genes have been previously reported to be clock controlled, heretofore there is no report about the circadian expression of 3 other genes. Considering the importance of oncogenes/TSGs in the initiation and progression of cancer, further studies are suggested for the identification of exact circadian expression patterns of these 3 human oncogenes/TSGs.
Key words
Oncogenes; tumor suppressor genes (TSGs); circadian expression; bioinformatic analysis